Neurodata Without Borders (NWB)
Type: Software,
Keywords: Neurophysiology, Data standard, FAIR data sharing, Intracellular electrophysiology, Extracellular electrophysiology, Optical physiology, Data science
Neurodata Without Borders (NWB) is a data standard for neurophysiology, providing neuroscientists with a common standard to share, archive, use, and build analysis tools for neurophysiology data.
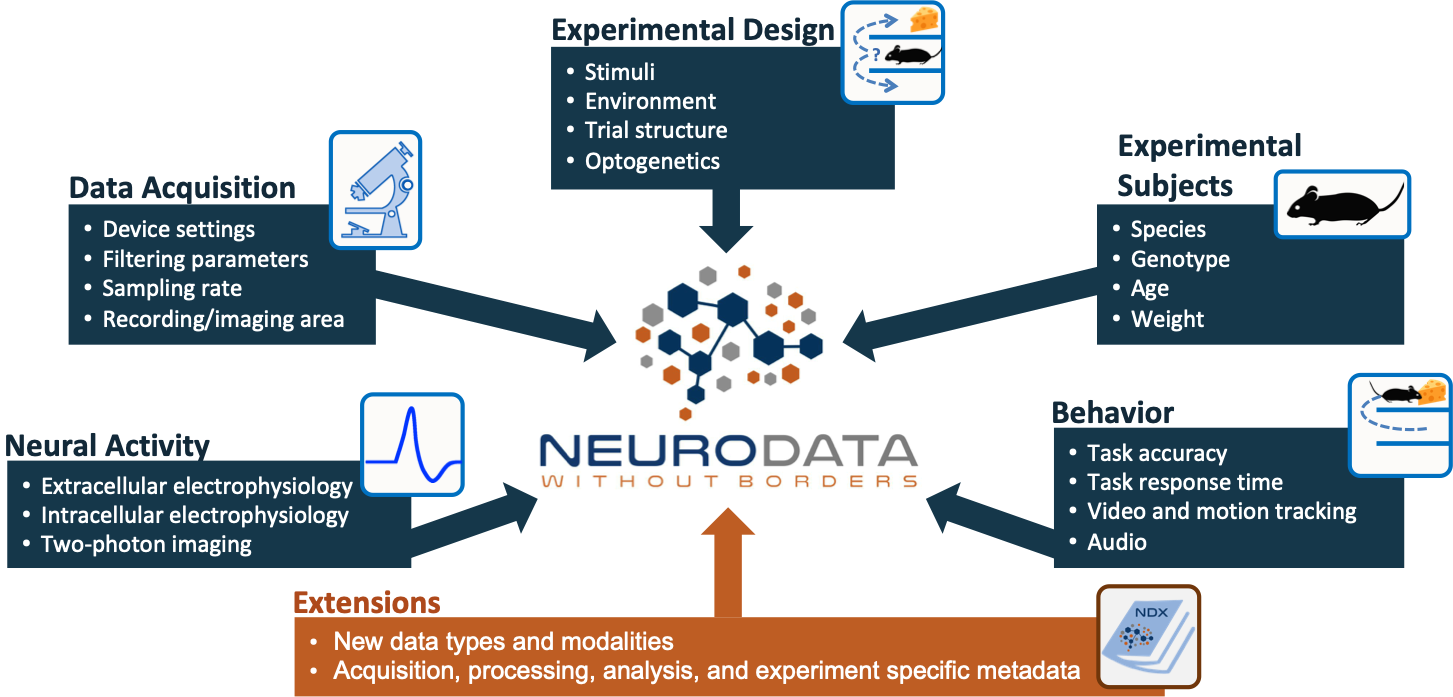
Neurodata Without Borders: Neurophysiology (NWB) is a data standard for neurophysiology, providing neuroscientists with a common standard to share, archive, use, and build common analysis tools for neurophysiology data. NWB is designed to store a variety of neurophysiology data, including data from intracellular and extracellular electrophysiology experiments, data from optical physiology experiments, and tracking and stimulus data. NWB is more than just a file format; it defines an ecosystem of tools, methods, and standards for storing, sharing, and analyzing complex neurophysiology data. NWB provides software for data standardization and application programming interfaces (APIs) for reading and writing the data, and is supported by a growing ecosystem of data analysis and management tools.
* Neurodata Without Borders (NWB) is a data standard for neurophysiology, providing neuroscientists with a common standard to share, archive, use, and build common analysis tools for neurophysiology data (see https://www.nwb.org/).
* NWB provides a comprehensive set of core tools for validating, creating, reading, and extending NWB data files and schema https://nwb-overview.readthedocs.io/en/latest/core_tools/core_tools_home.html
* NWB is supported by a growing ecosystem of neurophysiology tools, providing users access to state-of-the-art tools and methods for analyzing and sharing neurophysiology data (see https://nwb-overview.readthedocs.io/en/latest/tools/analysis_tools_home.html ).
* Publication and sharing of NWB neurophysiology data is supported by the DANDI BRAIN Initiative archive (see https://dandiarchive.org).
* INCF Endorsed Standard and Best Practice (see https://www.incf.org/sbp/nwb).
* 2019 R&D 100 Award Winner (see https://www.nwb.org/2019/10/31/neurodata-without-borders-wins-2019-rd100-award/).
* Store, share, analyze, publish, and reuse intracellular and extracellular electrophysiology and optical physiology data.
* Oliver Rübel, Andrew Tritt, Ryan Ly, Benjamin K. Dichter, Satrajit Ghosh, Lawrence Niu, Pamela Baker, Ivan Soltesz, Lydia Ng, Karel Svoboda, Loren Frank, Kristofer E. Bouchard, The Neurodata Without Borders ecosystem for neurophysiological data science, Oct. 2022, eLife 11:e78362. https://doi.org/10.7554/eLife.78362 (bioRxiv preprint: https://doi.org/10.1101/2021.03.13.435173)
* A. J. Tritt, O. Rübel, B. Dichter, R. Ly, D. Kang, E. F. Chang, L. M. Frank, K. E. Bouchard, “HDMF: Hierarchical Data Modeling Framework for Modern Science Data Standards,” IEEE International Conference on Big Data (Big Data), Los Angeles, CA, USA, December 2019, pp. 165-179. Online on IEEEXplore.
* Rübel, O., Tritt, A., Dichter, B., Braun, T., Cain, N., Clack, N., Davidson, T. J., Dougherty, M., Fillion-Robin, J.-C., Graddis, N., Grauer, M., Kiggins, J. T., Niu, L., Ozturk, D., Schroeder, W., Soltesz, I., Sommer, F. T., Svoboda, K., Ng, L., Frank, L. M., Bouchard, K. NWB:N 2.0: An Accessible Data Standard for Neurophysiology. January, 2019, doi: https://doi.org/10.1101/523035, bioRxiv preprint.
* Overview and getting started with NWB: https://nwb-overview.readthedocs.io/
* Online video training course: https://training.incf.org/collection/neurodata-without-borders-neurophysiology-nwbn
* NWB YouTube channel: https://www.youtube.com/c/NeurodataWithoutBorders/videos
* News and announcements: https://www.nwb.org/news/
* NWB Workshops: https://neurodatawithoutborders.github.io/nwb_hackathons/
NWB is supported by the DANDI BRAIN Initiative archive for publishing and sharing cellular neurophysiology data available at https://gui.dandiarchive.org.
Oliver Ruebel, Staff Scientists, Computational Research Division, Lawrence Berkeley National Laboratory; Benjamin Dichter, CatalystNeuro
Lawrence Berkeley National Laboratory, Berkeley, CA
TEAM / COLLABORATOR(S)
Ryan Ly, Computational Research Division, Lawrence Berkeley National Laboratory; Andrew Tritt, Computational Research Division, Lawrence Berkeley National Laboratory; Lawrence Niu, Vidrio Technologies
FUNDING SOURCE(S)
NIH U24NS120057

