MORF (MOnonucleotide Repeat Frameshift) Genetic Sparse Cell Labeling
Type: Molecular / Cellular,
Keywords: MORF, TIGRE-MORF (Ai166), Mouse, Morphology, Astrocyte, MORF3, Sparse labeling, Cre, Neuron, Microglia
Resource ID: RRID: SCR_021125
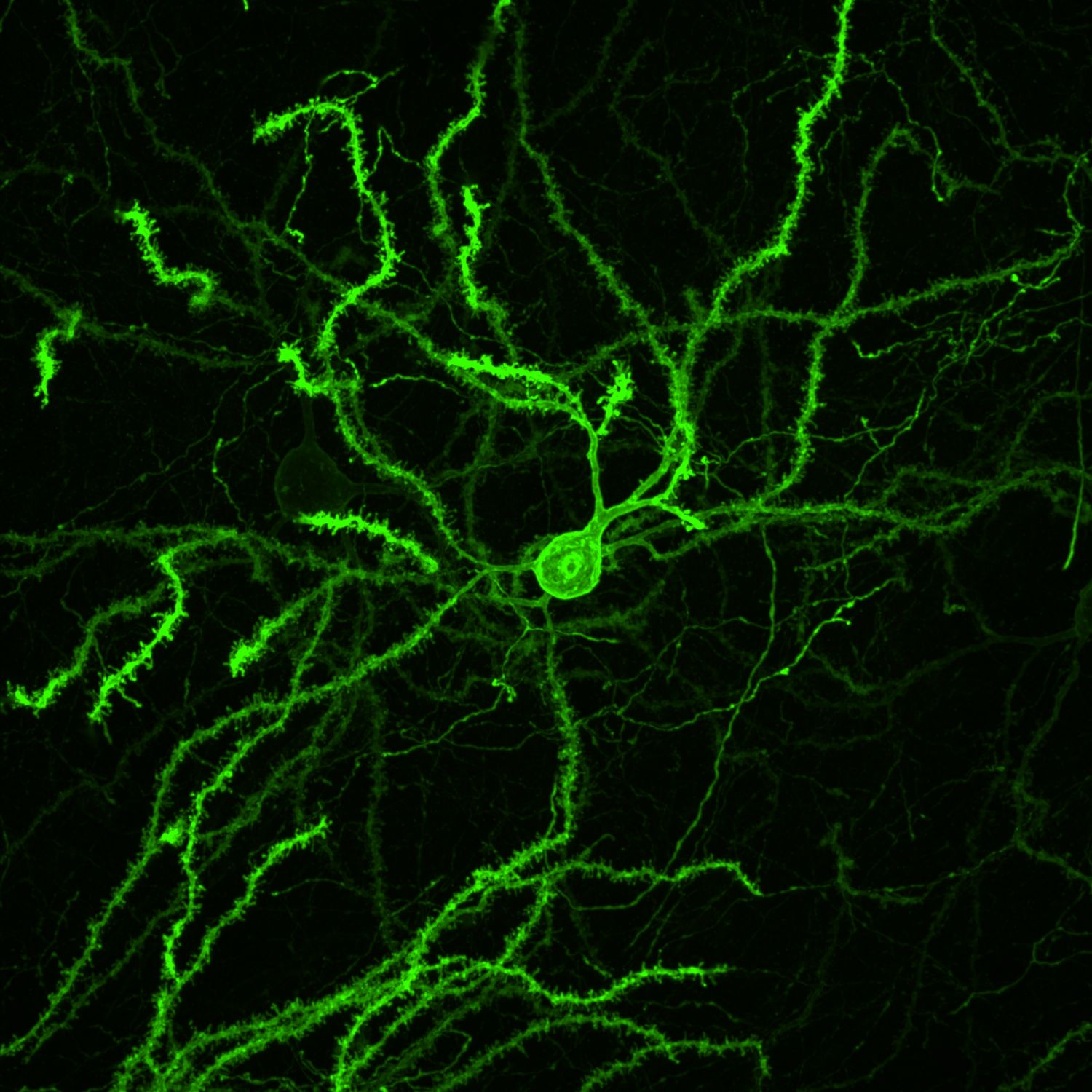
MORF mouse lines confer genetically-directed labeling of brain cell morphology
The MORF strategy addresses the need for a simple, generalizable, and scalable method to sparsely label genetically-defined neuronal populations by developing reporter mouse lines conferring Cre-dependent sparse cell labeling methodology based on MOnonucleotide Repeat Frameshift (MORF) as a stochastic translational switch. The suite of MORF reporter mice labels about 1-5% of Cre+ neurons and glia distributed stochastically throughout the brain and can be imaged with endogenous fluorescence (mNeonGreen in MORF1 and EGFP in TIGRE-MORF/Ai166) or stained for a multivalent immunoreporter (Spaghetti Monster fluorescent protein V5, or smFP-V5, in MORF3). As a resource for both the BRAIN Initiative and general neuroscience communities, MORF enables the labeling and reconstruction of thousands of genetically defined cells per brain for large-scale, unbiased classification and quantitative analyses of CNS cell types brainwide.
* MORF mouse lines include the strong, ubiquitous CAG promoter, LoxP-STOP-LoxP cassette for Cre-dependent expression, an unstable, out-of-frame mononucleotide repeat that can undergo frameshift that functions as translational switch for stochastic expression of a reporter gene.
* Different MORF lines make use of various membrane-bound reporters, including mNeonGreen-F (MORF1), EGFP-F (TIGER-MORF), spaghetti monster Fluorescent Protein V5 (smFP-V5-F; MORF3).
* MORF is compatible with various tissue processing (iDISCO+ and stochastic electrotransport clearing and immunolabeling) and imaging modalities (light microscopy, confocal, light sheet, electron microscopy).
* MORF3 is is the most versatile line with highest signal-to-noise, capable of spare labeling neurons and glial cells at 1-5% labeling frequency (less than 1% with inducible Cre lines).
The MORF reporters permit visualization of the complete morphology of genetically-defined neurons and glial cells at single cell resolution. By labeling cells at less than 1% of a given genetically-defined population with a membrane-bound reporter that has an extremely high signal-to-noise ratio, MORF allows the full 3D reconstruction and high-resolution analyses of the complete local morphology (dendrites) and far-reaching projections (axons) of CNS cell-types. The complete morphological visualization of MORF-labeled cells can be captured through various time-points in the lifespan of a given cell-type, from development through aging.
* Dendritic reconstruction of single neurons.
* Tracing and reconstruction of axonal projections of single neurons.
* Visualization and analysis of dendritic spines and axonal terminals at single neuron resolution.
*MORF1 mice (JAX: 035400)
*MORF3 mice (JAX: 035403)
*TIGRE-MORF or Ai166 mice (JAX: 035404)
* Genetically-defined, high-resolution single-cell imaging of CNS-cell types.
* A generalizable and scalable method that makes use of the existing repository of Cre-mouse lines.
* Incorporates well validated reporters that make MORF amenable to most tissue processing and imaging protocols.
*Veldman et al. 2020, Brainwide Genetic Sparse Cell Labeling to Illuminate the Morphology of Neurons and Glia with Cre-Dependent MORF Mice. Neuron vol. 108,1: 111-127.e6. doi:10.1016/j.neuron.2020.07.019
*Peng et al. 2021, Morphological diversity of single neurons in molecularly defined cell types. Nature vol. 598,7879 (2021): 174-181. doi:10.1038/s41586-021-03941-1
*Muñoz-Castañeda et al. 2021, Cellular anatomy of the mouse primary motor cortex. Nature vol. 598,7879: 159-166. doi:10.1038/s41586-021-03970-w
*BRAIN Initiative Cell Census Network (BICCN) 2021, A multimodal cell census and atlas of the mammalian primary motor cortex. Nature. 2021;598(7879):86-102. doi:10.1038/s41586-021-03950-0
X. William Yang, Principal Investigator
David Geffen School of Medicine, University of California, Los Angeles (UCLA)
TEAM / COLLABORATOR(S)
Matthew Veldman, Assistant Professor, Medical College of Wisconsin
Chang (Chris) Park, Project Scientist, University of California, Los Angeles (UCLA)
Hong-Wei Dong, Professor, University of California, Los Angeles (UCLA)
Hongkui Zeng, Executive Director, Allen Institute for Brain Science
FUNDING SOURCE(S)
NIH U01MH117079
NIH U01MH106008

